Rachel Melamed
Assistant Professor of Biology
University of Massachusetts at Lowell
Welcome
I an assistant professor at UMass Lowell in the Biology department, also affiliated with the Center of Biomedical and Health Research in Data Sciences. I’m excited to work with Biology students at all levels to make discoveries about the origins of disease.
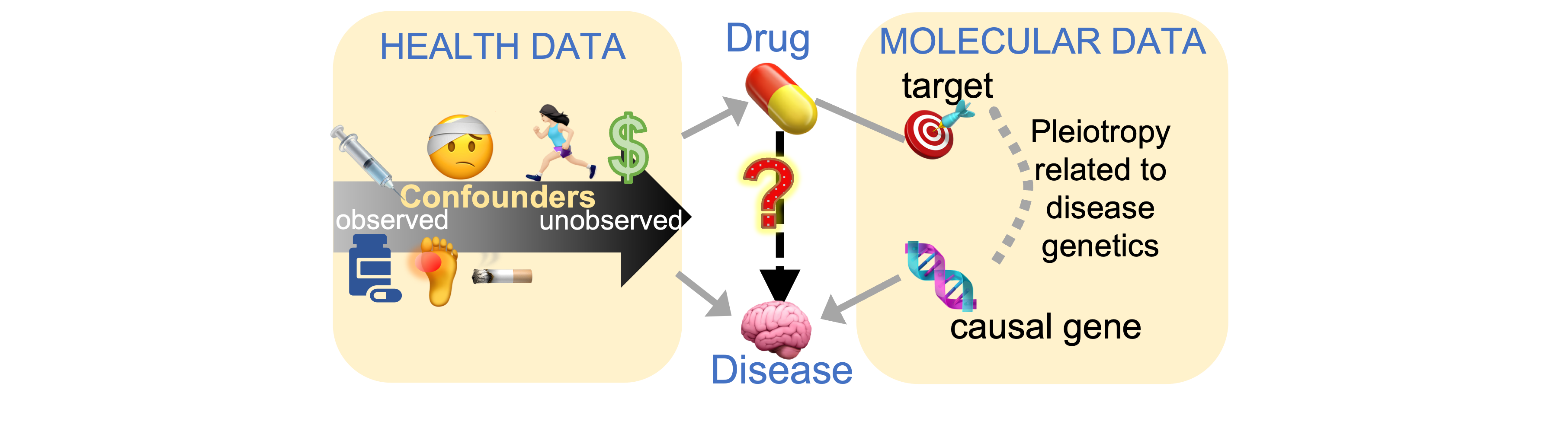
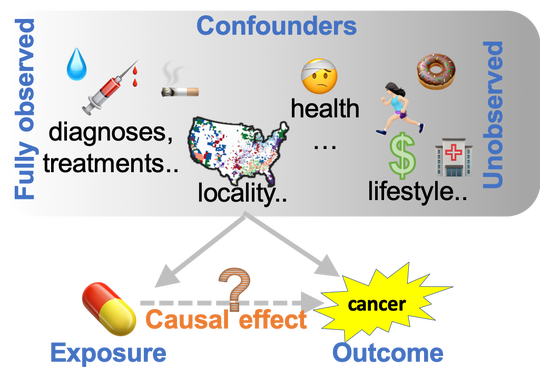
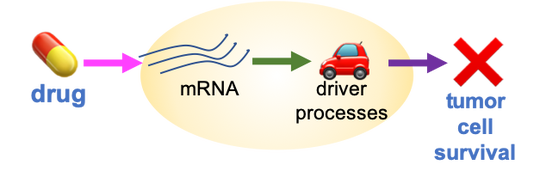
My goal is to discover health conditions, medical treatments, or other exposures that change risk of disease, by linking health data to molecular data. These results can uncover how the disease works and identify interventions to prevent and treat it.
To tackle this goal, our research brings together many kinds of data, including health records, biobanks, cancer genomics projects, and massive experimental studies of drugs. We will use methods inspired by computer science and epidemiology.
Interests
- Disease biology & disease genomics
- Data science
- Epidemiology & causal inference
Experience
Postdoc in Biomedical Data Science
University of Chicago
PhD in Biomedical Informatics
Columbia University
BA in Computer Science
Brown University